Dr Siddharth Srivastava, Chief Scientist (Molecular Oncology), Neuberg Center of Genomic Medicine elaborates on how genomics has changed the way cancer diagnosis and treatment is perceived
Precision medicine, personalised medicine, targeted drugs, next-generation sequencing and many more words are used frequently when people talk about cancer treatment. What exactly they mean and how DNA analysis of tumour is changing the way cancer is perceived and treated. To understand the whole story, let’s imagine a normal cell about to become cancerous. A normal cell multiples from one to two when it gets a growth signal usually from adjacent cells. These growth signals are small proteins that bind to the receptors on the surface of this particular cell. Upon binding to the receptor, the receptor protein changes its conformation and relays the signal to a specific signalling protein nearby. Let’s name it as signalling protein one. Signalling protein one triggers signalling protein two, which may trigger signalling protein three. The last signalling protein in the chain enters nucleus and sits over specific and marked portions of the DNA inside. This act allows specific genes in the nucleus to transcribe and translate new proteins which tell our cells to divide.
Thus, an external growth signal triggers a relay race of signalling proteins leading to growth and multiplication of cells. Off course this process must be tightly regulated else the cells would keep on dividing and end up in a large mass called tumour. Each of these signalling proteins are coded by genes. Genes are coded by four letters, A, T, G and C. Let’s imagine signalling protein two is coded by a ten letter long gene ATGCGATGCT. Unfortunately, our cell was residing in the lungs of a person who was a chain smoker. Through mechanisms unknown, the fourth letter of the gene changed from ‘C’ to a ‘T’. While this difference may seem very small, it made a big difference in the way signalling protein two behaved. Earlier signalling protein two needed a signal from signalling protein one to in turn trigger signalling protein three. However, with a change in the 4th letter, even without a signal from upstream, signalling protein two can trigger the protein downstream of it and that too, continuously.
Of course, the cell is constantly stimulated and it keeps dividing. In effect the cell has lost regulation over its own growth that leads to tumour formation. When a signalling protein gets mutated and causes cancer, they are called oncoprotein and the genes that were mutated are termed oncogenes. EGFR and KRAS are few such oncogenes and pathogenic mutations in those lead to NSCLC type of lung cancer.
On the other hand, a group of proteins act as guards to the process that allows the cell to proceed for cell cycle and eventually multiply from one cell into two. P53 is one such example. If a mutation occurs in the gene TP53 that codes for protein P53 such that now P53 is rendered useless, the cell will continuously go into cell cycle and multiply into a tumour. Such genes are called tumour suppressor genes. We have stated the mechanism or oncogenes and tumour suppressor genes in a simplistic fashion. More molecular mechanisms that may make a cell cancerous, exists. For e.g. aberrations in DNA repair.
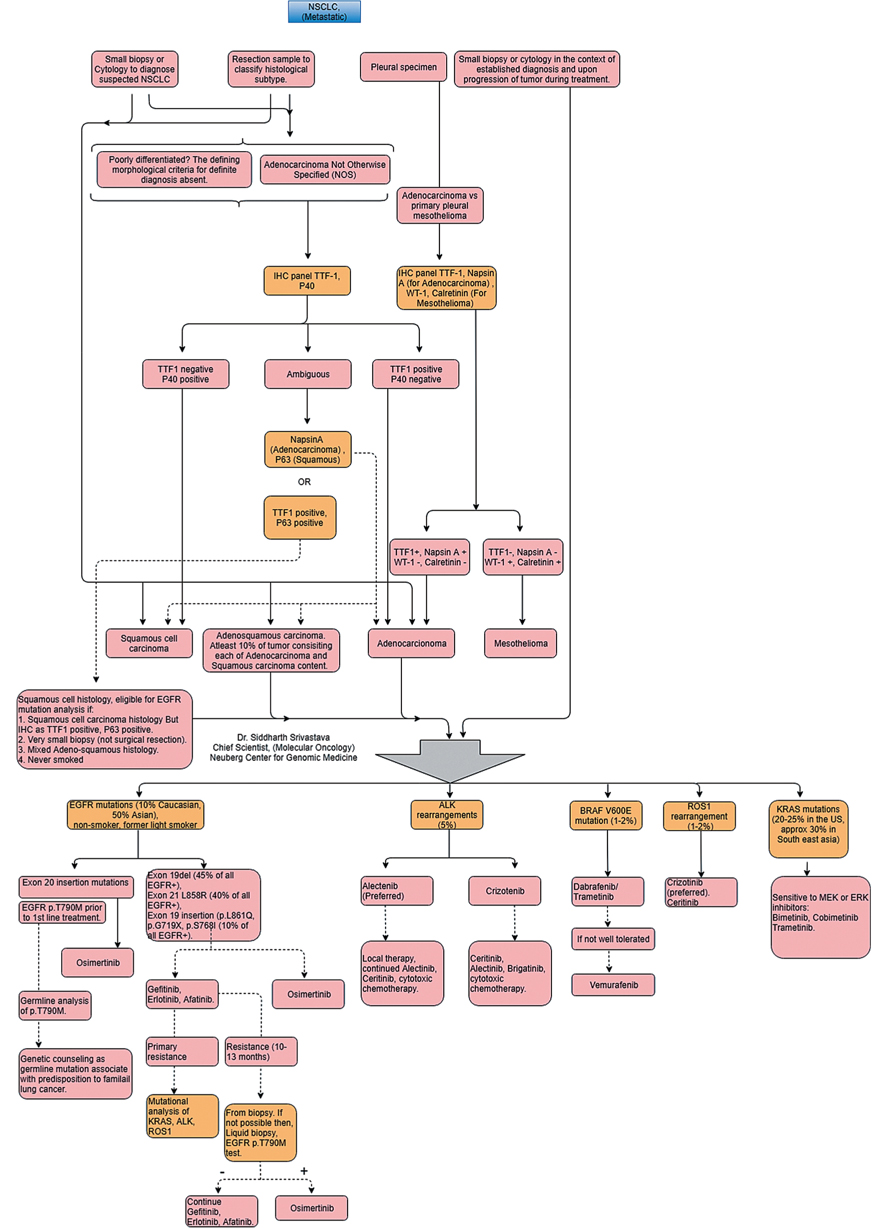
The knowledge that cancer is caused by specific mutations in genes led to large efforts to catalogue such mutations. The Cancer Genome Atlas https://cancergenome.nih.gov/, International Cancer Genome Consortium http://icgc.org/ are few examples. Such international efforts are describing landscape of mutations across the entire human genome and correlate them to specific cancer types. This knowledge also triggered efforts from biotechnology and pharmaceutical companies which are devising new molecules every month that targets oncoproteins.
These drugs are popularly called targeted therapies as the drugs target very specific protein molecules. Gefitinib (Iressa), Imatinib, Alectinib, Crizotinib are few examples. Let’s say if signalling protein two was mutated and caused continuous relaying to growth signal to cause cancer, oncologist would use a targeted therapy that specifically attacked and neutralise signalling protein two. However, such targeted therapy would not affect any other protein. Thus, to choose a targeted therapy, it is imperative to understand which protein is causing cancer and to do that, one needs to know which gene is mutated. Thus, unlike chemotherapy where much of the choice and regimen of treatment relied upon the clinical presentation and histopathology of the patient, targeted therapy could be chosen only if the oncologist knew which protein is causing cancer.
This opened entirely new disciplines of molecular diagnostics and molecular pathology. Molecular diagnostics deal with a gamut of technology focussed on extracting tumour DNA and reading it. Since the tissue samples are limiting, laboratories use PCR to amplify tiny DNA signals. These amplified DNA are then read using Sanger method, Next Generation Sequencing method or by droplet digital PCRs. Final goal of molecular diagnostics is to read the tumour DNA and compare it with healthy tissue.
The differences reveal critical characteristics of the cancer which is exploited by an oncologist to determine treatment and prognosis of the patient. The advent of molecular biology in cancer diagnosis and treatment has opened up requirement of molecular pathologists. Currently PhDs researching in pathology at molecular level, understand cancer pathology, cell biology and molecular biology. Based on molecular (DNA, RNA and proteins) differences between tumour cells and normal cells, molecular pathologists make recommendations to an oncologist in a fashion similar to a histo-pathologist.
References:
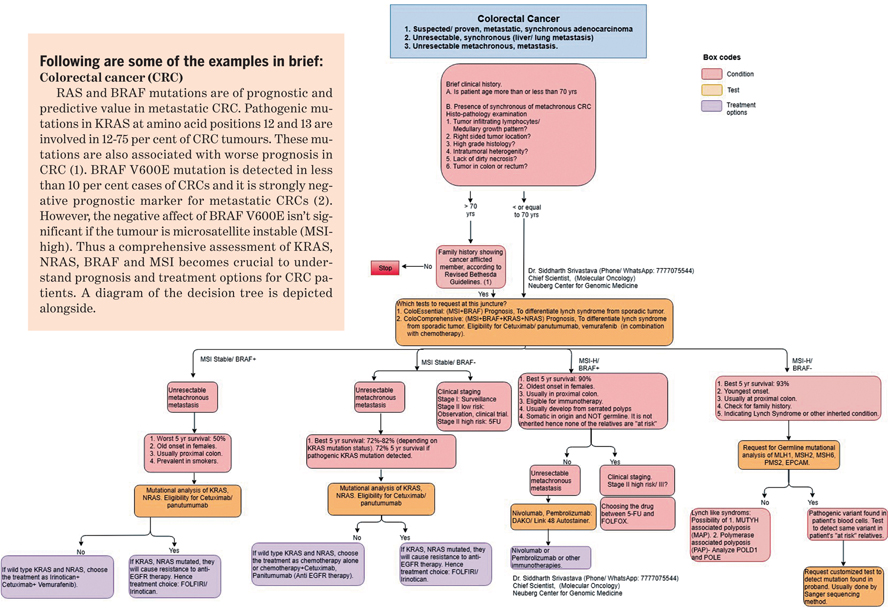
1. Porru M, Pompili L, Caruso C, Biroccio A, Leonetti C. Targeting KRAS in metastatic colorectal cancer: current strategies and emerging opportunities. J Exp Clin Cancer Res. 2018;37(1):57. Published 2018 Mar 13. doi:10.1186/s13046-018-0719-1
2. Ursem C, Atreya CE, Van Loon K. Emerging treatment options for BRAF-mutant colorectal cancer. Gastrointest Cancer. 2018;8:13–23. doi:10.2147/GICTT.S125940
- Advertisement -


